Mutants and controls
All IMPC statistical analyses compare observations in genetic mutant animals against observations of wildtype controls.
The IMPC protocols guarantee that all mutant measurements are accompanied by corresponding data from matched non-mutant animals, i.e. from animals assessed at the same time and at the same facility. As a result, the IMPC has a large collection of measurements pertaining to wildtype animals, all of which are annotated with metadata.
Although the wildtype data have some variability, as power analyses have shown, statistical tests benefit from using a wide set of control data points. Thus, unless the control metadata indicate substantial changes, mutant lines are compared against controls from their immediate control group as well as many other relevant animals. In practice, the effective control pool can include up to several thousand wildtype animals.
Changes in metadata that disqualify pooling controls include differences in location (mutants are compared only against wildtype animals processed at the same facility).
Since Data Release 15.1, the IMPC implements a soft windowing procedure which assigns more weight to the control mice collected proximally in time to the mutant mice.
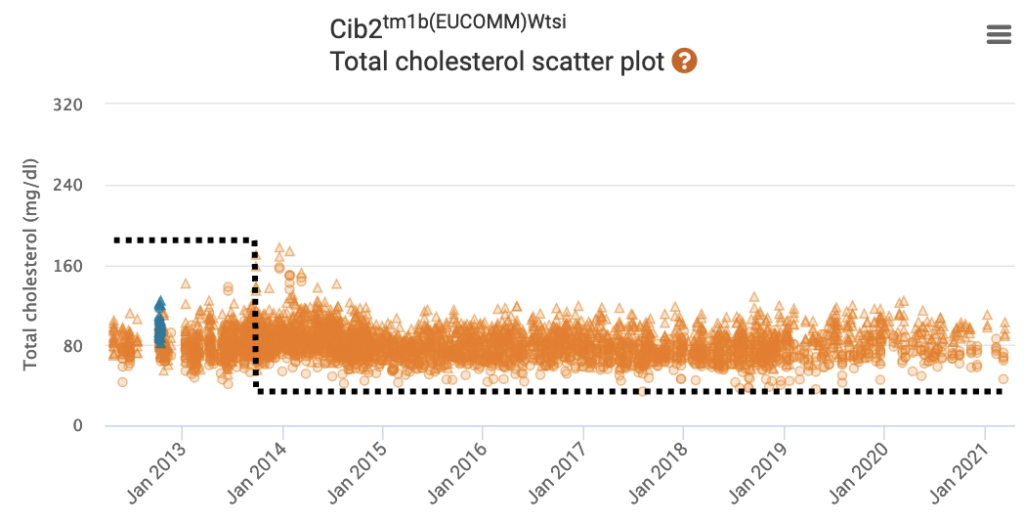
Learn more:
Read a News post about the soft windowing approach being implemented to data analysis in the IMPC website
Read the soft windowing paper: Soft Windowing Application to Improve Analysis of High-throughput Phenotyping Data, Journal of Bioinformatics 2019
Read about power analysis in this paper: Analysis of mammalian gene function through broad-based phenotypic screens across a consortium of mouse clinics, Nature Genetics 2015
